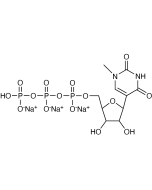
Pseudouridine-5'-Triphosphate - (N-1019)
Pseudouridine (5-ribosyluracil) was the first modified ribonucleoside discovered. It is the most abundant natural modified RNA base, and has been deemed the "fifth nucleoside" in RNA. It can be found in structural RNAs, such as transfer, ribosomal and small nuclear RNA. Pseudouridine has been found to enhance base stacking and translation. Pseudouridine-5'-triphosphate (Pseudo-UTP) is used to impart desirable mRNA characteristics such as increased nuclease stability, increased translation or altered interaction of innate immune receptors with in vitro transcribed RNA. Pseudo-UTP, along with 5-methylcytidine-5'-triphosphate (5-methyl-CTP) has shown innate immune suppression in culture and in vivo while enhancing translation in recent publications. For example, Warren et al. determined an efficient means of reprogramming multiple human cell types using modified mRNA that can express the four primary reprogramming proteins. These cells are referred to as induced pluripotency stem cells (iPSCs). Warren et al. found that enzymatically synthesized RNA substituted with Pseudo-UTP, 5-Methyl-CTP and ARCA effectively evaded the cell’s innate immune response, a crucial component in their success. Reduced toxicity due to substitution with Pseudo-UTP, 5-Methyl-CTP was critical since it allowed repeated transfection with in vitro transcribed mRNA over several weeks.
Katalin Karikó, prominent mRNA researcher, references TriLink as a leading manufacturer of Pseudo-UTP; "We value TriLink, it is an excellent company indeed. The product is very good and the customer service is very good too... everybody I know who is making pseudoU RNA is getting it from you."
| Catalog No | N-1019 |
|---|---|
| Purity | ≥97% by AX-HPLC |
| Extinction Coefficient | 7,546 Lmol-1cm-1 at 262 nm |
| Molecular Formula | C9H15N2O15P3 (free acid) |
| Molecular Weight | 484.10 g/mole (free acid) |
| Salt Form | Li+ |
| Concentration | 100 mM |
| Buffer | H2O |
| Recommended Storage | -20°C or below |
| Other Name(s) | Pseudo-UTP, 5-Ribosyl Uracil |
| Application | Aptamers, Epigenetics/DNA Damage, In vitro Transcription, Mutagenesis, Photocrosslinking Studies |
| Backbone | 5'-Triphosphate |
| Base Analog(s) | Pseudouridine |
| Sugar Type(s) | RNA |
| Nucleotide Category | Base Modified RNA |
N-1019 Safety Data Sheet
Ask an Expert
CoA search tool
Products are for research use only, not for use in diagnostic or therapeutic procedures or for use in humans. Products are not for resale without express written permission from TriLink No license under any patent or other intellectual property right of TriLink or its licensors is granted or implied by the purchase unless otherwise provided in writing.
TriLink does not warrant that the use or sale of the products delivered hereunder will not infringe the claims of any United States or other patents or patents pending covering the use of the product alone or in combination with other products or in the operation of any process. All and any use of TriLink product is the purchaser's sole responsibility.
- Madore, E.; Florentz, C.; Giegé, R.; Sekine, S.; Yokoyama, S.; Lapointe, J. . Effect of modified nucleotides on Escherichia coli tRNAGlu structure and on its aminoacylation by glutamyl-tRNA synthetase. Predominant and distinct roles of the mnm5 and s2 modifications of U34.
- Karikó, Katalin; Muramatsu, Hiromi; Welsh, Frank A.; Ludwig, János; Kato, Hiroki; Akira, Shizuo; Weissman, Drew . Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability.
- Karikó, Katalin; Buckstein, Michael; Ni, Houping; Weissman, Drew . Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA.
- Anderson, Bart R.; Muramatsu, Hiromi; Nallagatla, Subba R.; Bevilacqua, Philip C.; Sansing, Lauren H.; Weissman, Drew; Karikó, Katalin . Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation.
- Anderson, Bart R.; Muramatsu, Hiromi; Jha, Babal K.; Silverman, Robert H.; Weissman, Drew; Karikó, Katalin . Nucleoside modifications in RNA limit activation of 2'-5'-oligoadenylate synthetase and increase resistance to cleavage by RNase L.
- Karikó, Katalin; Muramatsu, Hiromi; Ludwig, János; Weissman, Drew . Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA.
- Anderson, B. R.; Karikó, K.; Weissman, D. . Nucleofection induces transient eIF2α phosphorylation by GCN2 and PERK.
- Karikó, Katalin; Muramatsu, Hiromi; Keller, Jason M.; Weissman, Drew . Increased erythropoiesis in mice injected with submicrogram quantities of pseudouridine-containing mRNA encoding erythropoietin.
- Charette, M.; Gray, M. W. . Pseudouridine in RNA: what, where, how, and why.
- Green, Nathaniel M.; Moody, Krishna-Sulayman; Debatis, Michelle; Marshak-Rothstein, Ann . Activation of autoreactive B cells by endogenous TLR7 and TLR3 RNA ligands.
- Warren, Luigi; Ni, Yuhui; Wang, Jiwu; Guo, Xirong . Feeder-free derivation of human induced pluripotent stem cells with messenger RNA.
- Uchida, Satoshi; Itaka, Keiji; Uchida, Hirokuni; Hayakawa, Kentaro; Ogata, Toru; Ishii, Takehiko; Fukushima, Shigeto; Osada, Kensuke; Kataoka, Kazunori . In vivo messenger RNA introduction into the central nervous system using polyplex nanomicelle.
- Guo, Xing Rong; Wang, Xiao Li; Li, Man Chol; Yuan, Ya Hong; Chen, Yun; Zou, Dan Dan; Bian, Liu Jiao; Li, Dong Sheng . PDX-1 mRNA-induced reprogramming of mouse pancreas-derived mesenchymal stem cells into insulin-producing cells in vitro.
- Wang, Xiao Li; Hu, Pei; Guo, Xing Rong; Yan, Ding; Yuan, Yahong; Yan, Shi Rong; Li, Dong Sheng . Reprogramming human umbilical cord mesenchymal stromal cells to islet-like cells with the use of in vitro–synthesized pancreatic-duodenal homebox 1 messenger RNA
- Baba, Miyuki; Itaka, Keiji; Kondo, Kenji; Yamasoba, Tatsuya; Kataoka, Kazunori . Treatment of neurological disorders by introducing mRNA in vivo using polyplex nanomicelles.
- Hausburg, Frauke; Na, Silke; Voronina, Natalia; Skorska, Anna; Müller, Paula; Steinhoff, Gustav; David, Robert . Defining optimized properties of modified mRNA to enhance virus- and DNA- independent protein expression in adult stem cells and fibroblasts.
- Lee, Kunwoo; Yu, Pengzhi; Lingampalli, Nithya; Kim, Hyun Jin; Tang, Richard; Murthy, Niren . Peptide-enhanced mRNA transfection in cultured mouse cardiac fibroblasts and direct reprogramming towards cardiomyocyte-like cells.
- Michel, Tatjana; Kankura, Anna; Salinas Medina, Martha L.; Kurz, Julia; Behring, Andreas; Avci-Adali, Meltem; Nolte, Andrea; Schlensak, Christian; Wendel, Hans Peter; Krajewski, Stefanie . In Vitro Evaluation of a Novel mRNA-Based Therapeutic Strategy for the Treatment of Patients Suffering from Alpha-1-Antitrypsin Deficiency.
- Pratico, Elizabeth D.; Feger, Bryan J.; Watson, Michael J.; Sullenger, Bruce A.; Bowles, Dawn E.; Milano, Carmelo A.; Nair, Smita K. . RNA-Mediated Reprogramming of Primary Adult Human Dermal Fibroblasts into c-kit(+) Cardiac Progenitor Cells.
- Uchida, Satoshi; Kataoka, Kazunori; Itaka, Keiji . Screening of mRNA Chemical Modification to Maximize Protein Expression with Reduced Immunogenicity.
- Wroblewska, Liliana; Kitada, Tasuku; Endo, Kei; Siciliano, Velia; Stillo, Breanna; Saito, Hirohide; Weiss, Ron . Mammalian synthetic circuits with RNA binding proteins for RNA-only delivery.
- Andries, Oliwia; Mc Cafferty, Séan; De Smedt, Stefaan C.; Weiss, Ron; Sanders, Niek N.; Kitada, Tasuku . N(1)-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice.
- Poleganov, Marco Alexander; Eminli, Sarah; Beissert, Tim; Herz, Stephanie; Moon, Jung-Il; Goldmann, Johanna; Beyer, Arianne; Heck, Rosario; Burkhart, Isabell; Barea Roldan, Diana; Türeci, Özlem; Yi, Kevin; Hamilton, Brad; Sahin, Ugur . Efficient Reprogramming of Human Fibroblasts and Blood-Derived Endothelial Progenitor Cells Using Nonmodified RNA for Reprogramming and Immune Evasion.
- Abraham, Meike-Kristin; Nolte, Andrea; Reus, Rebekka; Behring, Andreas; Zengerle, Diane; Avci-Adali, Meltem; Hohmann, Jan David; Peter, Karlheinz; Schlensak, Christian; Wendel, Hans Peter; Krajewski, Stefanie . In vitro Study of a Novel Stent Coating Using Modified CD39 Messenger RNA to Potentially Reduce Stent Angioplasty-Associated Complications.
- Endo, Kei; Hayashi, Karin; Saito, Hirohide . High-resolution Identification and Separation of Living Cell Types by Multiple microRNA-responsive Synthetic mRNAs.
- Koblas, Tomas; Leontovyc, Ivan; Loukotova, Sarka; Kosinova, Lucie; Saudek, Frantisek . Reprogramming of Pancreatic Exocrine Cells AR42J Into Insulin-producing Cells Using mRNAs for Pdx1, Ngn3, and MafA Transcription Factors.
- Dykstra, Brad; Lee, Jungmin; Mortensen, Luke J.; Yu, Haixiao; Wu, Zhengliang L.; Lin, Charles P.; Rossi, Derrick J.; Sackstein, Robert . Glycoengineering of E-Selectin Ligands by Intracellular versus Extracellular Fucosylation Differentially Affects Osteotropism of Human Mesenchymal Stem Cells.
- Kauffman, Kevin J.; Mir, Faryal F.; Jhunjhunwala, Siddharth; Kaczmarek, James C.; Hurtado, Juan E.; Yang, Jung H.; Webber, Matthew J.; Kowalski, Piotr S.; Heartlein, Michael W.; DeRosa, Frank; Anderson, Daniel G. . Efficacy and immunogenicity of unmodified and pseudouridine-modified mRNA delivered systemically with lipid nanoparticles in vivo.
- Goparaju, Sravan Kumar; Kohda, Kazuhisa; Ibata, Keiji; Soma, Atsumi; Nakatake, Yukhi; Akiyama, Tomohiko; Wakabayashi, Shunichi; Matsushita, Misako; Sakota, Miki; Kimura, Hiromi; Yuzaki, Michisuke; Ko, Shigeru B. H.; Ko, Minoru S. H. . Rapid differentiation of human pluripotent stem cells into functional neurons by mRNAs encoding transcription factors.
- Guo, Xing-Rong; Yang, Zhuo-Shun; Tang, Xiang-Jun; Zou, Dan-Dan; Gui, Hui; Wang, Xiao-Li; Ma, Shi-Nan; Yuan, Ya-Hong; Fang, Juan; Wang, Bin; Zhang, Li; Sun, Xu-Yong; Warnock, Garth L.; Dai, Long-Jun; Tu, Han-Jun . The application of mRNA-based gene transfer in mesenchymal stem cell-mediated cytotoxicity of glioma cells.
- Michel, Tatjana; Luft, Daniel; Abraham, Meike-Kristin; Reinhardt, Sabrina; Salinas Medina, Martha L.; Kurz, Julia; Schaller, Martin; Avci-Adali, Meltem; Schlensak, Christian; Peter, Karlheinz; Wendel, Hans Peter; Wang, Xiaowei; Krajewski, Stefanie . Cationic Nanoliposomes Meet mRNA: Efficient Delivery of Modified mRNA Using Hemocompatible and Stable Vectors for Therapeutic Applications.
- Leontovyc, Ivan; Habart, David; Loukotova, Sarka; Kosinova, Lucie; Kriz, Jan; Saudek, Frantisek; Koblas, Tomas . Synthetic mRNA is a more reliable tool for the delivery of DNA-targeting proteins into the cell nucleus than fusion with a protein transduction domain.
- Chen, Jiahuan; Fu, Yi; Day, Daniel S.; Sun, Ye; Wang, Shiyan; Liang, Xiaodong; Gu, Fei; Zhang, Fang; Stevens, Sean M.; Zhou, Pingzhu; Li, Kai; Zhang, Yan; Lin, Ruei-Zeng; Smith, Lois E. H.; Zhang, Jin; Sun, Kun; Melero-Martin, Juan M.; Han, Zeguang; Park, Peter J.; Zhang, Bing; Pu, William T. . VEGF amplifies transcription through ETS1 acetylation to enable angiogenesis.
- Shibata, Tomonori; Fujita, Yoshihiko; Ohno, Hirohisa; Suzuki, Yuki; Hayashi, Karin; Komatsu, Kaoru R.; Kawasaki, Shunsuke; Hidaka, Kumi; Yonehara, Shin; Sugiyama, Hiroshi; Endo, Masayuki; Saito, Hirohide . Protein-driven RNA nanostructured devices that function in vitro and control mammalian cell fate.
- Oh, Sanders; Kessler, John A. . Design, Assembly, Production, and Transfection of Synthetic Modified mRNA.
- Wang, Yuhua; Zhang, Lu; Xu, Zhenghong; Miao, Lei; Huang, Leaf . mRNA Vaccine with Antigen-Specific Checkpoint Blockade Induces an Enhanced Immune Response against Established Melanoma.
- Liu, Lina; Wang, Yuhua; Miao, Lei; Liu, Qi; Musetti, Sara; Li, Jun; Huang, Leaf . Combination Immunotherapy of MUC1 mRNA Nano-vaccine and CTLA-4 Blockade Effectively Inhibits Growth of Triple Negative Breast Cancer.
- Kogut, Igor; McCarthy, Sandra M.; Pavlova, Maryna; Astling, David P.; Chen, Xiaomi; Jakimenko, Ana; Jones, Kenneth L.; Getahun, Andrew; Cambier, John C.; Pasmooij, Anna M. G.; Jonkman, Marcel F.; Roop, Dennis R.; Bilousova, Ganna . High-efficiency RNA-based reprogramming of human primary fibroblasts.
- Suknuntha, Kran; Tao, Lihong; Brok-Volchanskaya, Vera; D'Souza, Saritha S.; Kumar, Akhilesh; Slukvin, Igor . Optimization of Synthetic mRNA for Highly Efficient Translation and its Application in the Generation of Endothelial and Hematopoietic Cells from Human and Primate Pluripotent Stem Cells.
- Potapov, Vladimir; Fu, Xiaoqing; Dai, Nan; Corrêa, Ivan R. Jr; Tanner, Nathan A.; Ong, Jennifer L. . Base modifications affecting RNA polymerase and reverse transcriptase fidelity.
- Golombek, Sonia; Pilz, Martin; Steinle, Heidrun; Kochba, Efrat; Levin, Yotam; Lunter, Dominique; Schlensak, Christian; Wendel, Hans Peter; Avci-Adali, Meltem . Intradermal Delivery of Synthetic mRNA Using Hollow Microneedles for Efficient and Rapid Production of Exogenous Proteins in Skin.
- Mondal, Nandini; Dykstra, Brad; Lee, Jungmin; Ashline, David J.; Reinhold, Vernon N.; Rossi, Derrick J.; Sackstein, Robert . Distinct human α(1,3)-fucosyltransferases drive Lewis-X/sialyl Lewis-X assembly in human cells.
- Steinle, Heidrun; Ionescu, Tudor-Mihai; Schenk, Selina; Golombek, Sonia; Kunnakattu, Silju-John; Özbek, Melek Tutku; Schlensak, Christian; Wendel, Hans Peter; Avci-Adali, Meltem . Incorporation of Synthetic mRNA in Injectable Chitosan-Alginate Hybrid Hydrogels for Local and Sustained Expression of Exogenous Proteins in Cells.
- Kim, Bo-Eun; Choi, Soon Won; Shin, Ji-Hee; Kim, Jae-Jun; Kang, Insung; Lee, Byung-Chul; Lee, Jin Young; Kook, Myoung Geun; Kang, Kyung-Sun . Single-Factor SOX2 Mediates Direct Neural Reprogramming of Human Mesenchymal Stem Cells via Transfection of In Vitro Transcribed mRNA.
- Heinz, Sven; Texari, Lorane; Hayes, Michael G. B.; Urbanowski, Matthew; Chang, Max W.; Givarkes, Ninvita; Rialdi, Alexander; White, Kris M.; Albrecht, Randy A.; Pache, Lars; Marazzi, Ivan; García-Sastre, Adolfo; Shaw, Megan L.; Benner, Christopher . Transcription Elongation Can Affect Genome 3D Structure.
- Loomis, Kristin H.; Lindsay, Kevin E.; Zurla, Chiara; Bhosle, Sushma M.; Vanover, Daryll A.; Blanchard, Emmeline L.; Kirschman, Jonathan L.; Bellamkonda, Ravi V.; Santangelo, Philip J. . In Vitro Transcribed mRNA Vaccines with Programmable Stimulation of Innate Immunity.
- Pardi, N;Hogan, MJ;Pelc, RS;Muramatsu, H;Andersen, H;DeMaso, CR;Dowd, KA;Sutherland, LL;Scearce, RM;Parks, R;Wagner, W;Granados, A;Greenhouse, J;Walker, M;Willis, E;Yu, JS;McGee, CE;Sempowski, GD;Mui, BL;Tam, YK;Huang, YJ;Vanlandingham, D;Holmes, VM;Balachandran, H;Sahu, S;Lifton, M;Higgs, S;Hensley, SE;Madden, TD;Hope, MJ;Karik . Zika virus protection by a single low-dose nucleoside-modified mRNA vaccination
- Zhao, W;Zeng, C;Yan, J;Du, S;Hou, X;Zhang, C;Li, W;Deng, B;McComb, DW;Xue, Y;Kang, DD;Dong, Y; . Construction of Messenger RNA (mRNA) Probes Delivered By Lipid Nanoparticles to Visualize Intracellular Protein Expression and Localization at Organelles
- Zhang, X;Zhao, W;Nguyen, G;Zhang, C;Zeng, C;Yan, J;Du, S;Hou, X;Li, W;Jiang, J;Deng, B;McComb, D;Dorkin, R;Shah, A;Barrera, L;Gregoire, F;Singh, M;Chen, D;Sabatino, D;Dong, Y; . Functionalized lipid-like nanoparticles for in vivo mRNA delivery and base editing
- Li, B;Zeng, C;Dong, Y; . Design and assessment of engineered CRISPR-Cpf1 and its use for genome editing
- Melo, M;Porter, E;Zhang, Y;Silva, M;Li, N;Dobosh, B;Liguori, A;Skog, P;Landais, E;Menis, S;Sok, D;Nemazee, D;Schief, W;Weiss, R;Irvine, D; . Immunogenicity of RNA replicons encoding HIV env immunogens designed for self-assembly into nanoparticles
- Nance, K;Gamage, S;Alam, M;Yang, A;Levy, M;Link, C;Florens, L;Washburn, M;Gu, S;Oppenheim, J;Meier, J; . Cytidine acetylation yields a hypoinflammatory synthetic messenger RNA
- Wang, P;Logeart-Avramoglou, D;Petite, H;Goncalves, C;Midoux, P;Perche, F;Pichon, C; . Co-delivery of NS1 and BMP2 mRNAs to murine pluripotent stem cells leads to enhanced BMP-2 expression and osteogenic differentiation
- Lockhart, J;Canfield, J;Mong, EF;VanWye, J;Totary-Jain, H; . Nucleotide Modification Alters MicroRNA-Dependent Silencing of MicroRNA Switches
- Magadum, A;Singh, N;AnuKurian, A;KabirSharkar, M;Chepurko, E;Zangi, L; . Ablation of a single N-glycosylation site in human FSTL1 induces cardiomyocyte proliferation and cardiac regeneration
- Michel, T;Luft, D;Abraham, MK;Reinhardt, S;Salinas Medina, ML;Kurz, J;Schaller, M;Avci-Adali, M;Schlensak, C;Peter, K;Wendel, HP;Wang, X;Krajewski, S; . Cationic nanoliposomes meet mRNA: Efficient delivery of modified mRNA using hemocompatible and stable vectors for therapeutic applications
- Zhuang, X;Qi, Y;Wang, M;Yu, N;Nan, F;Zhang, H;Tian, M;Li, C;Lu, H;Jin, N; . mRNA Vaccines Encoding the HA Protein of Influenza A H1N1 Virus Delivered by Cationic Lipid Nanoparticles Induce Protective Immune Responses in Mice
- Zhang, M;Sun, J;Li, M;Jin, X; . Modified mRNA-LNP vaccines confer protection against experimental DENV-2 infection in mice
- Sadegh, C;Ebina, W;Arvanites, AC;Davidow, LS;Rubin, LL;Macklis, JD; . Synthetic modified Fezf2 mRNA (modRNA) with concurrent small molecule SIRT1 inhibition enhances refinement of cortical subcerebral/corticospinal neuron identity from mouse embryonic stem cells
- Leppek, K;Byeon, GW;Kladwang, W;Wayment-Steele, HK;Kerr, CH;Xu, AF;Kim, DS;Topkar, VV;Choe, C;Rothschild, D;Tiu, GC;Wellington-Oguri, R;Fujii, K;Sharma, E;Watkins, AM;Nicol, JJ;Romano, J;Tunguz, B;Participants, E;Barna, M;Das, R; . Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
- Majumder, A;Suknuntha, K;Bennin, D;Klemm, L;Brok-Volchanskaya, V;Huttenlocher, A;Slukvin, I; . Generation of Human Neutrophils from Induced Pluripotent Stem Cells in Chemically Defined Conditions Using ETV2 Modified mRNA
- Hirosawa, M;Saito, H; . Cell-Type-Specific CRISPR-Cas9 System with miRNAs
- Zhang, W; . Investigation and Characterization of RNA Modifications
- Abraham, MK;Peter, K;Michel, T;Wendel, HP;Krajewski, S;Wang, X; . Nanoliposomes for Safe and Efficient Therapeutic mRNA Delivery: A Step Toward Nanotheranostics in Inflammatory and Cardiovascular Diseases as well as Cancer
- Svitkin, YV;Gingras, AC;Sonenberg, N; . Membrane-dependent relief of translation elongation
- Lin, X;Chen, H;Xie, Y;Zhou, X;Wang, Y;Zhou, J;Long, S;Hu, Z;Zhang, S;Qiu, W;Zeng, Z;Liu, L; . Combination of CTLA-4 blockade with MUC1 mRNA nanovaccine induces enhanced anti-tumor CTL activity by modulating tumor microenvironment of triple negative breast cancer
- Fujita, Y;Hirosawa, M;Hayashi, K;Hatani, T;Yoshida, Y;Yamamoto, T;Saito, H; . A versatile and robust cell purification system with an RNA-only circuit composed of microRNA-responsive ON and OFF switches
- Evers, MJW;Du, W;Yang, Q;Kooijmans, SAA;Vink, A;van Steenbergen, M;Vader, P;de Jager, SCA;Fuchs, SA;Mastrobattista, E;Sluijter, JPG;Lei, Z;Schiffelers, R; . Delivery of modified mRNA to damaged myocardium by systemic administration of lipid nanoparticles
- Valentini, P;Pierattini, B;Zacco, E;Mangoni, D;Espinoza, S;Webster, N;Andrews, B;Carninci, P;Tartaglia, G;Pandolfini, L;Gustincich, S; . Towards SINEUP-based therapeutics: design of an in vitro synthesized SINEUP RNA
- Tang, X;Peng, H;Xu, P;Zhang, L;Fu, R;Tu, H;Guo, X;Huang, K;Lu, J;Chen, H;Dong, Z;Dai, L;Luo, J;Chen, Q; . Synthetic mRNA-based gene therapy for glioblastoma: TRAIL mRNA synergistically enhance PTEN mRNA-based therapy
- Peng, H;Guo, X;He, J;Duan, C;Yang, M;Zhang, X;Zhang, L;Fu, R;Wang, B;Wang, D;Chen, H;Xie, M;Feng, P;Dai, L;Tang, X;Luo, J; . Intracranial delivery of synthetic mRNA to suppress glioblastoma
- Jensen, TI;Mikkelsen, NS;Gao, Z;Fo . Targeted regulation of transcription in primary cells using CRISPRa and CRISPRi
- Yang, J;Ding, S; . Chimeric RNA binding protein based killing switch targeting hepatocellular carcinoma cells